BioByte 146: How Pleiades Predicts Alzheimer's, New EDEN Models Enable Drug Design via Evolutionary Datasets, Exploring the Inverse Relationship between Peripheral Cancer and AD, and Synthetic Ecology
Welcome to Decoding Bio’s BioByte: each week our writing collective highlight notable news—from the latest scientific papers to the latest funding rounds—and everything in between. All in one place.
What we read
Blogs
Using Interpretability to Identify a Novel Class of Alzheimer’s Biomarkers [Wang et al., Goodfire, January 2026]
Why it matters: Researchers from Goodfire AI utilize mechanistic interpretability methods to uncover the features that Pleiades, an epigenetics foundation model, uses to successfully predict Alzheimer’s Disease (AD) from cell-free DNA (cfDNA) samples.
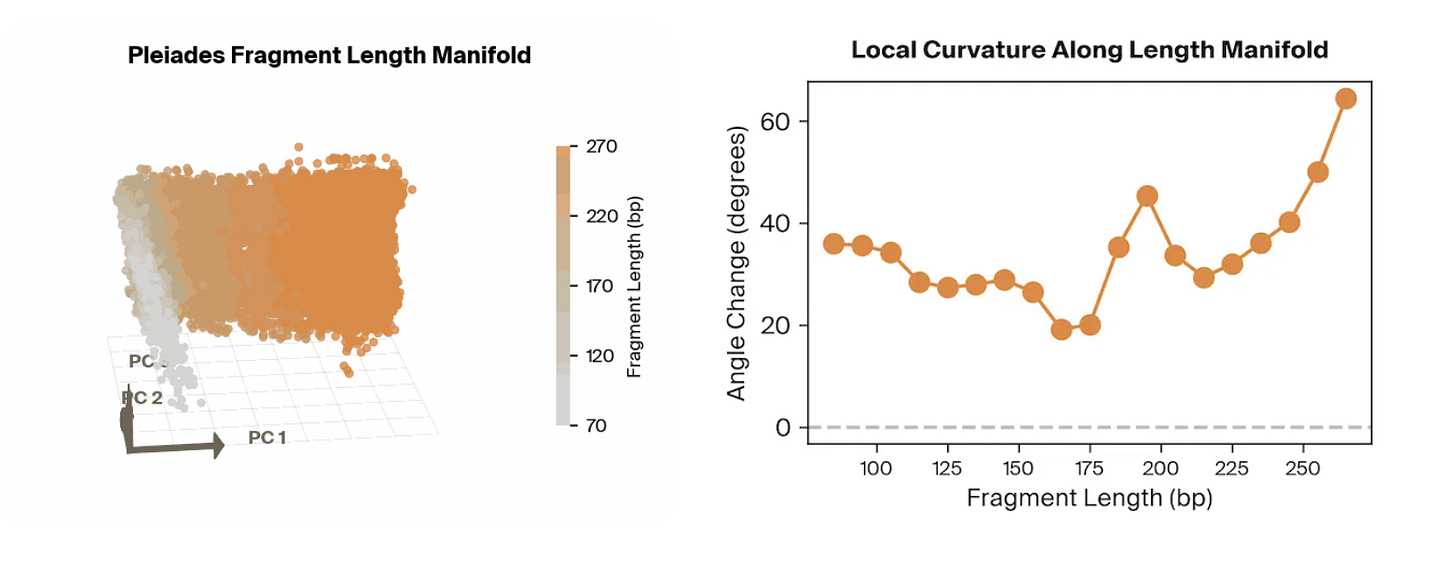
Pleiades is an epigenomic foundation model developed by Prima Mente that captures methylated and unmethylated human DNA from full genomes and cfDNA. They utilize a hierarchical set-attention architecture that enables them to expand attention across a large swath of DNA, such as for pools of cfDNA. In the original preprint, one powerful application Pleiades was pointed towards was the prediction of whether or not a patient had AD based on their cfDNA sequences, which it did extremely well in with an AUROC of 0.82.
The team at Goodfire applied mechanistic interpretability techniques to the frozen, cfDNA-fine-tuned Pleiades model, probing its final sample-level representations to identify what features enabled it to effectively predict AD status. Rather than relying on post-hoc feature attribution on raw inputs, they decomposed the internal embedding space using linear probes and sparse autoencoders to recover what qualities the embeddings contained.
Surprisingly, the strongest features utilized by the model were all correlated to the length of the input cfDNA sequences. With a simple classifier that takes in fragment length features alone, they achieved an AUROC of approximately 0.78 on an independent cohort. By combining fragment length with other biologically meaningful signals identified through their probing, such as methylation- and cell-type–related features, they achieved performance around 0.84 AUROC, approaching the full Pleiades cfDNA-only performance of ~0.82–0.83. More broadly, this work demonstrates how mechanistic interpretability can turn opaque foundation-model representations into concrete, testable biological hypotheses, helping bridge the gap between high-performing black-box predictors and mechanistic understanding of the disease signals they exploit.
Papers
Peripheral cancer attenuates amyloid pathology in Alzheimer’s disease via cystatin-c activation of TREM2 [Li et al., Cell, January 2026]
...Why it matters: Alzheimer’s disease therapies have largely focused on limiting the formation of amyloid plaques. However, once plaques are
This excerpt is provided for preview purposes. Full article content is available on the original publication.